文章目录
练习与作业1:作图
用下面的数据作图
- 利用下面代码读取一个样本的宏基因组相对丰度数据
abu <-
read_delim(
file = "../data/talk06/relative_abundance_for_RUN_ERR1072629_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");- 取前5个丰度最高的菌,将其它的相对丰度相加并归为一类
Qita; - 用得到的数据画如下的空心pie chart:
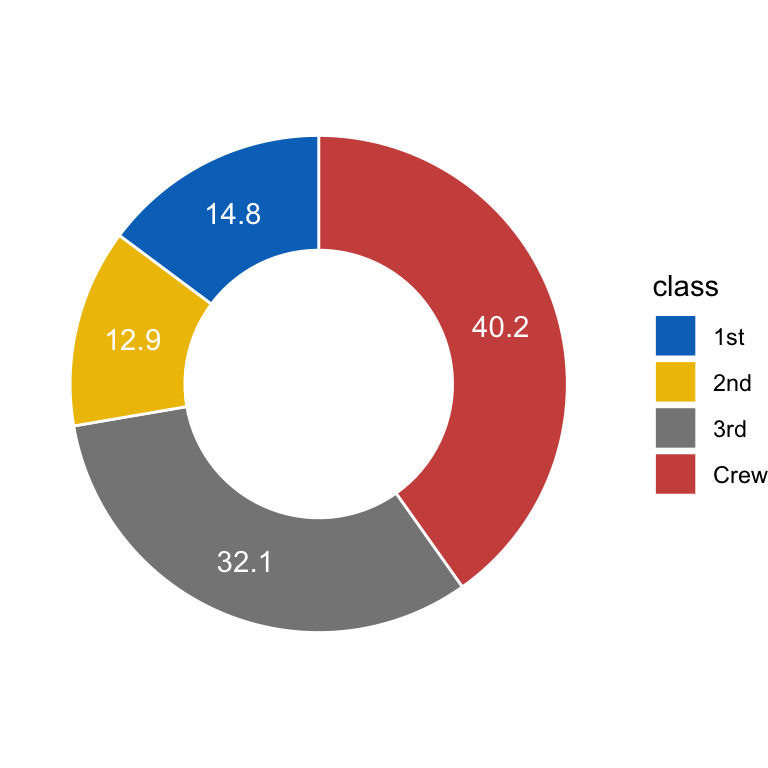
```{r}
## 代码写这里,并运行;
require(tidytidbits)
require(tidyverse)
abu <-
read_delim(
file = "../data/talk06/relative_abundance_for_RUN_ERR1072629_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");
toppart <- abu %>% arrange(-relative_abundance)%>%
lump_rows (scientific_name,relative_abundance,n=5,other_level="Qita")
ggplot(toppart, aes(x= 2, y = relative_abundance, fill = scientific_name)) +
geom_col() +
geom_text(aes(label = relative_abundance),
position = position_stack(vjust =0.5)) +
coord_polar(theta = "y") +
xlim(c(0.2, 3.5))+
theme_void()
```使用starwars变量做图
- 统计
starwars中hair_color的种类与人数时,可用下面的代码:
但是,怎么做到按数量从小到大排序?
```{r}
library(dplyr)
library(ggplot2)
library(forcats)
ggplot(starwars, aes(x = hair_color)) +
geom_bar() +
coord_flip()
``````{r}
## 代码写这里,并运行;
starshair <- starwars %>% group_by(hair_color) %>%
summarise(number=n_distinct(name))
starshair
ggplot(starshair,aes(x=fct_reorder(hair_color,number,.desc=T),y=number))+
geom_bar(stat = "identity")+
coord_flip()+xlab("hair_color") + ylab( "count" )
```统计 skin_color 时,将出现频率小于0.05(即5%)的颜色归为一类Others,按出现次数排序后,做与上面类似的 barplot;
```{r}
## 代码写这里,并运行;
starskin <- starwars %>% group_by(skin_color) %>%
summarise(count=n_distinct(name),
apparence=count/(n_distinct(starwars$name)))%>%
arrange(-count)%>%
lump_rows(skin_color,count,n=6,other_level = "Others")
ggplot(starskin,aes(x=fct_reorder(skin_color,count,.desc = T),y=count))+
geom_bar(stat = "identity")+
coord_flip()+xlab("skin_color")+ylab("count")
```使用 2 的统计结果,但画图时,调整 bar 的顺序,使得 Others 处于第4的位置上。提示,可使用 fct_relevel 函数;
```{r}
## 代码写这里,并运行;
ggplot(starskin,aes(x=fct_reorder(skin_color,count,.desc = T)%>%
fct_relevel("Others",after = 3),y=count))+
geom_bar(stat = "identity")+
coord_flip()+xlab("skin_color")+ylab("count")
```练习与作业2:数据分析
使用 STRING PPI 数据分析并作图
- 使用以下代码,装入PPI数据;
ppi <- read_delim( file = "../data/talk06/ppi900.txt.gz", col_names = T,
delim = "\t", quote = "" );- 随机挑选一个基因,得到类似于本章第一部分的互作网络图;
```{r}
## 代码写这里,并运行;
ppi <- read_delim( file = "../data/talk06/ppi900.txt.gz", col_names = T,
delim = "\t", quote = "" );
targetgene <-sample(ppi$gene1,1)
toppart <- ppi %>% filter( gene1 == targetgene ) %>%
arrange( desc( score ) ) %>% slice( 1:10 );
genes <- unique( c( targetgene, toppart$gene2 ) );
netdata <- ppi %>% filter( gene1 %in% genes & gene2 %in% genes );
if (!require("igraph")){
chooseCRANmirror();
install.packages("igraph");
}
library( igraph );
netdata.nr <-
netdata %>%
mutate( group =
if_else( gene1 > gene2,
str_c( gene1, gene2, sep = "-" ),
str_c( gene2, gene1, sep = "-" ) ) ) %>%
group_by( group ) %>% slice( 1 );
netnet.nr <- graph_from_data_frame( netdata.nr, directed = FALSE );
plot(netnet.nr);
```对宏基因组 相对丰度数据 进行分析
1.data/talk06 目录下有6个文本文件,每个包含了一个宏基因组样本的分析结果:
relative_abundance_for_curated_sample_PRJEB6070-DE-073_at_taxonlevel_species.txt
relative_abundance_for_curated_sample_PRJEB6070-DE-074_at_taxonlevel_species.txt
relative_abundance_for_curated_sample_PRJEB6070-DE-075_at_taxonlevel_species.txt
relative_abundance_for_curated_sample_PRJEB6070-DE-076_at_taxonlevel_species.txt
relative_abundance_for_curated_sample_PRJEB6070-DE-077_at_taxonlevel_species.txt- 分别读取以上文件,提取
scientific_name和relative_abundance两列; - 添加一列为样本名,比如
PRJEB6070-DE-073,PRJEB6070-DE-074… ; - 以
scientific_name为key,将其内容合并为一个data.frame或tibble,其中每行为一个样本,每列为样本的物种相对丰度。注意:用join或者spread都可以,只要能解决问题。 - 将
NA值改为0。
```{r}
## 代码写这里,并运行;
genesample1 <- read_delim(
file = "data/talk06/relative_abundance_for_curated_sample_PRJEB6070-DE-073_at_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");
genesample1 <- genesample1 %>% select(scientific_name=scientific_name,
relativa_abundance=taxon_rank_level)
genesample1 <-add_column(genesample1,samplename=c("PRJEB6070-DE-073"))
genesample2 <- read_delim(
file = "data/talk06/relative_abundance_for_curated_sample_PRJEB6070-DE-074_at_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");
genesample2 <- genesample2 %>% select(scientific_name=scientific_name,
relativa_abundance=taxon_rank_level)
genesample2 <-add_column(genesample2,samplename=c("PRJEB6070-DE-074"))
genesample3 <- read_delim(
file = "data/talk06/relative_abundance_for_curated_sample_PRJEB6070-DE-075_at_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");
genesample3 <- genesample3 %>% select(scientific_name=scientific_name,
relativa_abundance=taxon_rank_level)
genesample3 <-add_column(genesample3,samplename=c("PRJEB6070-DE-075"))
genesample4 <- read_delim(
file = "data/talk06/relative_abundance_for_curated_sample_PRJEB6070-DE-076_at_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");
genesample4 <- genesample4 %>% select(scientific_name=scientific_name,
relativa_abundance=taxon_rank_level)
genesample4 <- genesample4[-90,]
genesample4 <-add_column(genesample4,samplename=c("PRJEB6070-DE-076"))
genesample5 <- read_delim(
file = "data/talk06/relative_abundance_for_curated_sample_PRJEB6070-DE-077_at_taxonlevel_species.txt",
delim = "\t", quote = "", comment = "#");
genesample5 <- genesample5 %>% select(scientific_name=scientific_name,
relativa_abundance=taxon_rank_level)
genesample5 <- genesample5[-75,]
genesample5 <-add_column(genesample5,samplename=c("PRJEB6070-DE-077"))
genesample1_1 <- genesample1 %>% spread(scientific_name,relativa_abundance)
genesample2_1 <- genesample2 %>% spread(scientific_name,relativa_abundance)
genesample3_1 <- genesample3 %>% spread(scientific_name,relativa_abundance)
genesample4_1 <- genesample4 %>% spread(scientific_name,relativa_abundance)
genesample5_1 <- genesample5 %>% spread(scientific_name,relativa_abundance)
comb <- bind_rows( genesample1_1, genesample2_1 )%>%bind_rows(genesample3_1)%>%
bind_rows(genesample4_1)%>%bind_rows(genesample5_1)
comb[is.na(comb)]<-0
comb
```
COMMENTS | NOTHING